-
Faculty of Arts and HumanitiesJakobi 2, r 116-121 51005 Tartu linn, Tartu linn, Tartumaa ESTJakobi 2 51005 Tartu linn, Tartu linn, Tartumaa ESTJakobi 2, IV korrus 51005 Tartu linn, Tartu linn, Tartumaa ESTJakobi 2, III korrus, ruumid 302-337 51005 Tartu linn, Tartu linn, Tartumaa ESTÜlikooli 16 51003 Tartu linn, Tartu linn, Tartumaa ESTLossi 3 51003 Tartu linn, Tartu linn, Tartumaa ESTÜlikooli 18 50090 Tartu linn, Tartu linn, Tartumaa ESTPosti 1 71004 Viljandi linn, Viljandimaa ESTJakobi 2 51005 Tartu linn, Tartu linn, Tartumaa ESTJakobi 2 51005 Tartu linn, Tartu linn, Tartumaa ESTFaculty of Social SciencesLossi 36 51003 Tartu linn, Tartu linn, Tartumaa ESTJakobi 5 51005 Tartu linn, Tartu linn, Tartumaa ESTLossi 36, ruum 301 51003 Tartu linn, Tartu linn, Tartumaa ESTNarva mnt 18 51009 Tartu linn, Tartu linn, Tartumaa ESTNäituse 2 50409 Tartu linn, Tartu linn, Tartumaa ESTNäituse 20 - 324 50409 Tartu linn, Tartu linn, Tartumaa ESTLossi 36 51003 Tartu linn, Tartu linn, Tartumaa ESTRaekoja plats 2 20307 Narva linn, Ida-Virumaa ESTRingi 35 80012 Pärnu linn, Pärnu linn, Pärnumaa ESTLossi 36 51003 Tartu linn, Tartu linn, Tartumaa ESTLossi 36 51003 Tartu linn, Tartu linn, Tartumaa ESTFaculty of MedicineRavila 19 50411 Tartu linn, Tartu linn, Tartumaa ESTBiomeedikum, Ravila 19 50411 Tartu linn, Tartu linn, Tartumaa ESTNooruse 1 50411 Tartu linn, Tartu linn, Tartumaa ESTL. Puusepa 1a 50406 Tartu linn, Tartu linn, Tartumaa ESTL. Puusepa 8 50406 Tartu linn, Tartu linn, Tartumaa ESTRavila 19 50411 Tartu linn, Tartu linn, Tartumaa ESTUjula 4 51008 Tartu linn, Tartu linn, Tartumaa ESTRavila 50411 Tartu linn, Tartu linn, Tartumaa ESTRavila 19 50411 Tartu linn, Tartu linn, Tartumaa ESTFaculty of Science and TechnologyVanemuise 46 - 208 51003 Tartu linn, Tartu linn, Tartumaa ESTNarva mnt 18 51009 Tartu linn, Tartu linn, Tartumaa ESTRiia 23b/2 51010 Tartu linn, Tartu linn, Tartumaa ESTRavila 14a 50411 Tartu linn, Tartu linn, Tartumaa ESTNarva mnt 18 51009 Tartu linn, Tartu linn, Tartumaa ESTRiia 23, 23b - 134 51010 Tartu linn, Tartu linn, Tartumaa ESTObservatooriumi 1 61602 Tõravere alevik, Nõo vald, Tartumaa ESTNooruse 1 50411 Tartu linn, Tartu linn, Tartumaa ESTJ. Liivi tn 2 50409 Tartu linn, Tartu linn, Tartumaa ESTVanemuise 46 51003 Tartu linn, Tartu linn, Tartumaa ESTVanemuise 46 51003 Tartu linn, Tartu linn, Tartumaa ESTArea of Academic SecretaryLossi 3 51003 Tartu linn, Tartu linn, Tartumaa ESTUppsala 6, Lossi 36 51003 Tartu linn, Tartu linn, Tartumaa ESTArea of Head of FinanceÜlikooli 17 51005 Tartu linn, Tartu linn, Tartumaa ESTArea of Director of AdministrationÜlikooli 18A (III korrus) 51005 Tartu linn, Tartu linn, Tartumaa ESTÜlikooli 18, ruumid 102, 104, 209, 210 50090 Tartu linn, Tartu linn, Tartumaa ESTArea of Vice Rector for ResearchW. Struve 1 50091 Tartu linn, Tartu linn, Tartumaa ESTArea of Vice Rector for DevelopmentNarva mnt 18 51009 Tartu linn, Tartu linn, Tartumaa ESTVanemuise 46 51003 Tartu linn, Tartu linn, Tartumaa ESTLossi 25 51003 Tartu linn, Tartu linn, Tartumaa ESTArea of RectorArea of Vice Rector for Academic AffairsUppsala 10 51003 Tartu linn, Tartu linn, Tartumaa ESTÜlikooli 18b 51005 Tartu linn, Tartu linn, Tartumaa EST
replication
transcription
modifications of histones
Chromatin research group
Survival and life of cells depend on accurate and faithful inheritance and expression of their genetic information in the course of DNA replication and gene transcription. These processes are highly regulated in multiple levels, including by the regulation of DNA accessibility in chromatin. The general aim of our research is to clarify the role of chromatin in regulation of these fundamental processes in eukaryotic cells.
We are interested in revealing the mechanisms how chromatin modifications support the correct initiation of DNA replication and gene transcription, and what is the role of different chromatin-interacting proteins in recruitment of DNA replication and transcription machinery to the correct initiation sites of these processes. In particular, we are focused on studying the proteins that modify histone proteins, or are important in detection of histone modifications in chromatin. We use budding yeast (Saccharomyces cerevisiae) as a model organism, which has several technical advantages like fast growth and availability of very broad variety of genetic and biochemical research methods and assays.
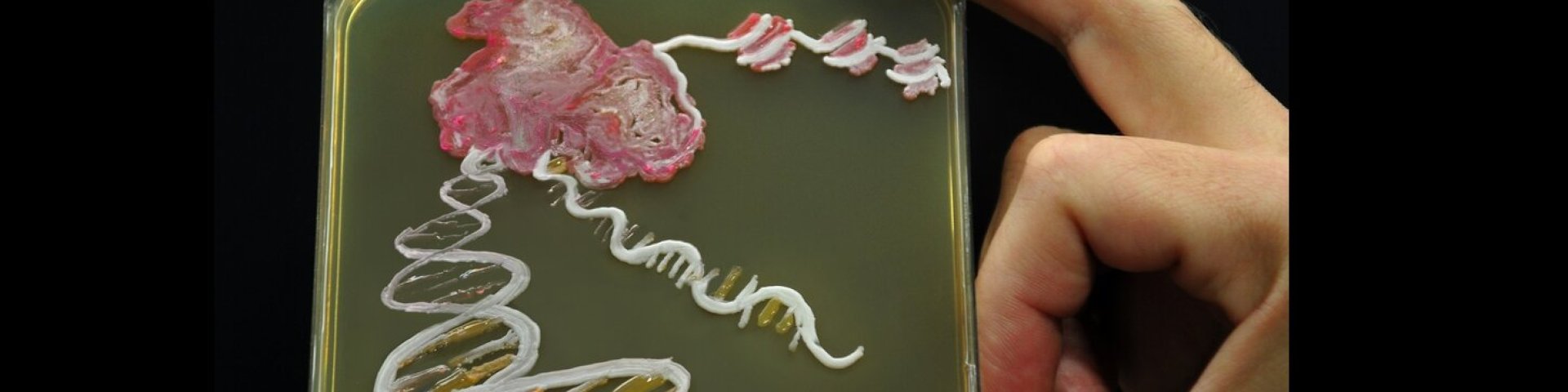
Header image: RNA polymerase gene transcribing (Image on a "painted" plate with yeast medium using different yeast species).
Idea and implementation: Mari Ann Rebane, Signe Värv, Sulev Kuuse, Arnold Kristjuhan
Reinapae, A., Ilves, I., Jürgens, H., Värv, S., Kristjuhan, K., & Kristjuhan, A. (2023). Interactions between Fkh1 monomers stabilize its binding to DNA replication origins. The Journal of biological chemistry, 299(8), 105026.
Peil, K., Jürgens, H., Luige, J., Kristjuhan, K., Kristjuhan, A. (2020). Taf14 is required for the stabilization of transcription pre‑initiation complex in Saccharomyces cerevisiae. Epigenetics & Chromatin 13: 24.
Sein, H., Reinmets, K., Peil, K., Kristjuhan, K., Värv, S., Kristjuhan, A. (2018). Rpb9-deficient cells are defective in DNA damage response and require histone H3 acetylation for survival. Scientific Reports 8: 2949.
Reinapae, A., Jalakas, K., Avvakumov, N., Lõoke, M., Kristjuhan, K., Kristjuhan, A. (2017). Recruitment of Fkh1 to replication origins requires precisely positioned Fkh1/2 binding sites and concurrent assembly of the prereplicative complex. PLoS Genetics 13: e1006588.
(in Estonian)
Novaator, 19.11.2024: Uudne meetod aitab leida kohalikust pärmirikkusest parimad töörügajad


