Ribosome, RNA modifications and protein synthesis research group
We are interested on the fundamental aspects of protein biosynthesis. In this respect we use E. coli as a model system. We study functional topography of the bacterial ribosome. In addition, we analyze ribosome biogenesis and ribosome degradation in bacteria. Currently our main research focus is the roles of modified nucleotides of the rRNA during ribosome functioning. More specifically, we analyze 23S rRNA modifications around the ribosomal peptidyltransferase center (PTC) in the domain V of 23S rRNA. 13 out of 25 modifications localize in this 23S rRNA region.
An E. coli strain lacking 12 modifications in the domain V of 23S rRNA was constructed. The resulting strain (del10, according to number of deleted genes) is barely viable at optimal growth conditions. Thus, effect of the rRNA modifications analyzed is additive. Remarkably, the rate of peptide bond formation on the del10 ribosomes is slower as compared to that of the wt ribosomes, demonstrating an importance of the rRNA modifications on the peptide bond synthesis on ribosomes. On the other hand, slow growth phenotype and other severe defects of del10 strain enable us to define functional roles of the individual modifications of rRNA by using addback strains. One of the genes, which is absent in del10 strain is reintroduced into the genome of del10. Phenotypic analysis of the addback strains has revealed unexpected new functions of the modified nucleotides during translation in vivo, bacterial susceptibility to antibiotics, cold sensitivity, and ribosome biogenesis. Effects of the modifications around the PTC on the ribosome structure is analyzed in cooperation with prof. M. Selmer’s group in Uppsala University.
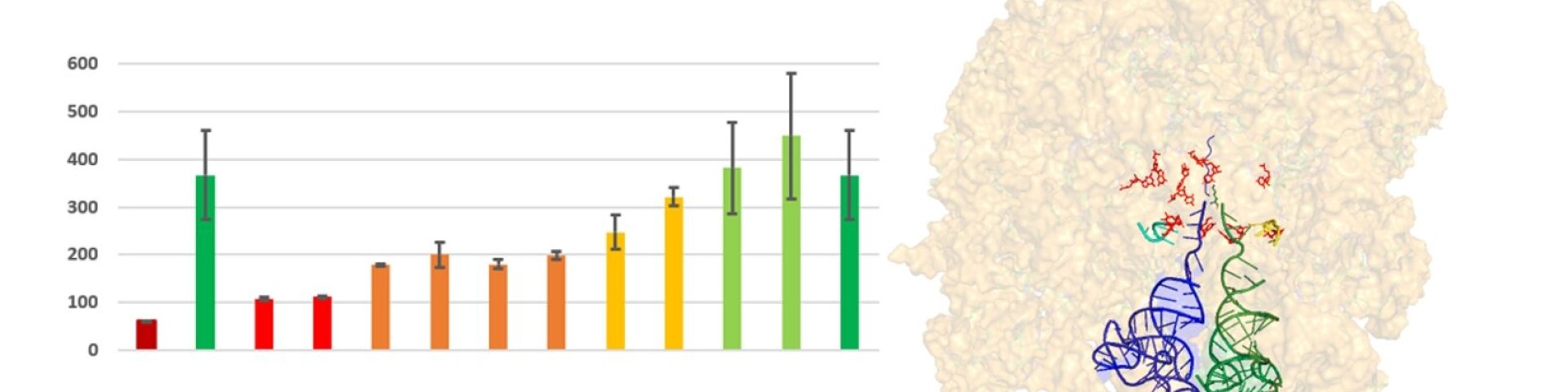
Scheme used in the header: Large subunit of ribosome showing A and P tRNAs and modified nucleotides in PTC and graph of recovery of deleted modifications at different bacterial growth rates (author: Jaanus Remme)
- Bao, L., Liljeruhm, J., Crespo Blanco, R., Brandis, G., Remme, J., Forster, A.C. Translational impacts of enzymes that modify ribosomal RNA around the peptidyl transferase centre. RNA Biol, 21(1):31-41
- Reier, K., Liiv, A., Remme, J. SILAC analysis of Escherichia coli proteome during progression of growth from exponential to prolonged stationary phase. Microbiol Resour Announc. 2024 Jun 11;13(6)
- Ero, R., Leppik, M., Reier, K., Liiv, A., Remme, J. (2024). Ribosomal RNA modification enzymes stimulate large ribosome subunit assembly in E. coli. Nucleic Acids Research, 2024, gkae222
- Reier, K., Liiv, A. and Remme, J. (2023) Ribosome Protein Composition Mediates Translation during the E. coli Stationary Phase. International Journal of Molecular Sciences 24, no. 4: 3128
- Jürgenstein, K., Tagel, M., Ilves, H., Leppik, M., Kivisaar, M., and Remme, J. (2022) Variance in translational fidelity of different bacterial species is affected by pseudouridines in the tRNA anticodon stem-loop. RNA Biology, 19, 1050-1058
- Reier, K., Lahtvee, P-J., Liiv, A., Remme, J. (2022) A Conundrum of r-Protein Stability: Unbalanced Stoichiometry of r-Proteins during Stationary Phase in Escherichia coli. mBio 13, (5) e01873-22
- Liljeruhm, J., Leppik, M., Bao, L., Truu, T., Noriega, M.C., Freyer, M.S., Liiv, A., Wang, J., Blanco, R.C., Ero, R., Remme, J., and Forster, A.C (2022) Plasticity and conditional essentiality of modification enzymes for domain V of Escherichia coli 23S ribosomal RNA. RNA 28, 796-807
- Datta, M., Pillai, M., Modak, M. J., Liiv, A., Khaja, F. T., Hussain, T., Remme, J., & Varshney, U. (2021). A mutation in the ribosomal protein uS12 reveals novel functions of its universally conserved PNSA loop. Molecular microbiology, 115(6), 1292–1308
(in Estonian)
Novaator: Kuidas panna kokku elus masin? Tartu Ülikooli teadlased teavad vastust
Uudishimu tippkeskus (ERR): Kuidas katku ajal paremat pidu pidada?
Postimees: Rekordkiired vaktsiinid on vaid murdosa uue biotehnoloogia lubadustest
Novaator: Vaktsiinist vähiravimini: TÜ teadlased katsetavad mRNA potentsiaali